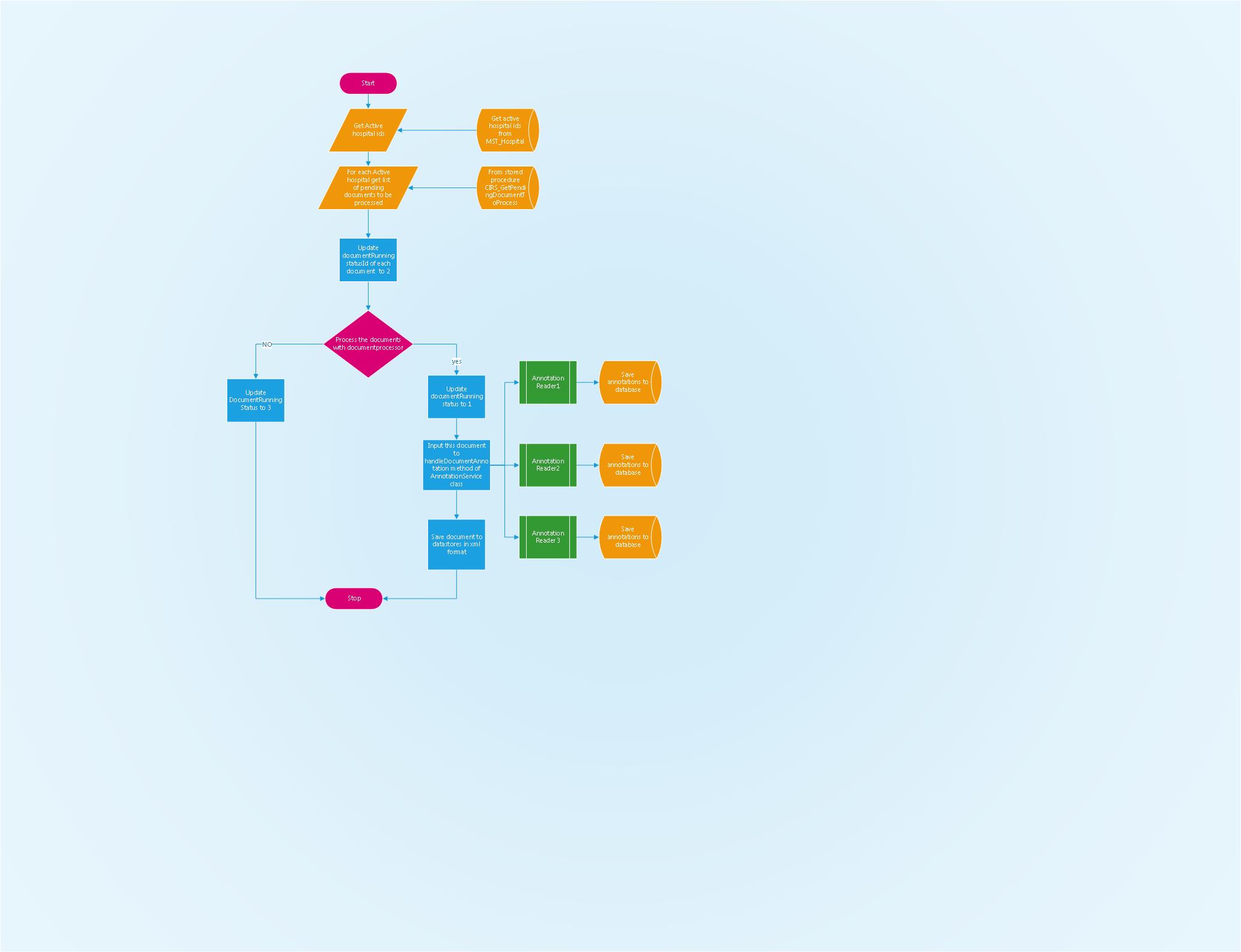
Workflow of cirs-gapp
Getting active hospitalids from database
AS our application supports multiple hospitals, we get the ids of the hospitals whose active status="1" in the database from a table called "MST_Hospital"
Getting documents that are to be processed
After getting ids of active hospitals from database , then we will get all the pending documents that are to be processed for each active hospital by using the stored procedure "CIRS_GetPendingDocumentToProcess"
| select Top 50 hdd.DocumentID,1 hospitalId,hdd.Document,hd.DocumentTypeID,mdt.DocumentAliasName,hd.TranscriptionDateTime,hd.AccountNumber,
hv.PatientClass,DATEDIFF(MONTH,hp.DOB,hv.AdmitDate)/12 as Age,DATEDIFF(DAY,hp.DOB,hv.AdmitDate) as AgeInDays,hp.Gender,hd.HospitalDay,
ISNULL(CONVERT(VARCHAR(10),hv.AdmitDate,101),CONVERT(VARCHAR(10),getDate(),101)) AdmitDate,ISNULL(CONVERT(VARCHAR(10),hv.DischargeDate,101),CONVERT(VARCHAR(10),getDate(),101)) DischargeDate,
CONVERT(VARCHAR(10),hp.DOB,101) DateOfBirth,idt.InternalDocumentTypeID,idt.InternalDocumentName
from HIM_DocumentData hdd
inner join HIM_Document hd on hdd.DocumentID=hd.DocumentID
inner join MST_DocumentType mdt on mdt.DocumentTypeID=hd.DocumentTypeID
inner join MST_InternalDocumentType idt on mdt.InternalDocumentTypeID=idt.InternalDocumentTypeID
inner join HIM_Visit hv on hv.AccountNumber=hd.AccountNumber
inner join HIM_Patient hp on hp.PatientId=hv.PatientID
where hd.DocumentRunningStatusID=0 and hd.Formatted=1 and hd.HospitalDay is not null
|
here we will connect to each hospital database by using the jdbcTemplate which is generated dynamically based on the id of the hospital.
After getting the documents that are to be processed from the database , we create gate documents from these documents by appending different features along with dictionary running features and jape running features,which will be required while processing the document through the gapp file
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23 | for(ClinicalDocument clinicalDocument: clinicalDocuments) {
Document gateDocument=Factory.newDocument(clinicalDocument.getDocument());
FeatureMap featureMap=Factory.newFeatureMap();
featureMap.put("documentId", clinicalDocument.getDocumentId());
featureMap.put("accountNumber", clinicalDocument.getAccountNumber());
featureMap.put("documentName", clinicalDocument.getDocumentName());
featureMap.put("documentTypeId", clinicalDocument.getDocumentTypeId());
featureMap.put("documentAliasName",clinicalDocument.getDocumentAliasName());
featureMap.put("internalDocumentAliasName", clinicalDocument.getInternalDocumentAliasName());
featureMap.put("hospitalDay", clinicalDocument.getHospitalDay());
featureMap.put("worktype", clinicalDocument.getDocumentAliasName());
featureMap.put("ageInDays", clinicalDocument.getAgeInDays());
featureMap.put("age", clinicalDocument.getAge());
featureMap.put("admitDate", clinicalDocument.getAdmitDate());
featureMap.put("dischargeDate", clinicalDocument.getDischargeDate());
featureMap.put("dateOfBirth", clinicalDocument.getDateOfBirth());
featureMap.put("gender", clinicalDocument.getGender());
featureMap.put("internalDocumentName",clinicalDocument.getInternalDocumentName());
clinicalDocumentRunningFeatures.setDictionaryRunningFeatures(featureMap);
clinicalDocumentRunningFeatures.setJapeRunningFeatures(featureMap, clinicalDocument.getInternalDocumentTypeId());
gateDocument.setFeatures(featureMap);
gateDocuments.add(gateDocument);
}
|
Processing the documents
Now these gate documents are ready to be processed ,we will pass these documents one by one to the processDocument method of the document Processor instance which we got from the gate-beans.xml
Before passing the documents to the processDocument method we update the documentRunningStatus of the document to 2 in the database.
If the document is processed successfully we will update the documentRunningStatus to 1 in the database.
Reading Annotations from document
Later these processed documents are passed to the handleDocumentAnnotations method of AnnotationService class , in which we will call different types of annotation readers to extract all the annotations from the processed document and save them to the database.we will discuss later in detail about the annotationService class and the different type of annotationReaders
Saving these analysed documents to xml
After analysis these documents are saved into the datastores in xml format based on the worktype and processing date of the document .These files when again loaded into gate developer will be helpful in displaying the annotations and their covered texts and also can be useful in debugging the missed annotations.
If any exception occurs in the above whole process then the documentrunningstatus will be updated to 3 in the database
Scheduling the application
All the above processes are called in a single method called processClinicalDocumentFromQueue, which is scheduled for every thirty seconds by using @Scheduled(fixedRate=time stamp) above the method signature which is achieved by enabling scheduling option in the main class by using @EnableScheduling annotation
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41 | @Scheduled(fixedRate= 30000)
public void processClinicalDocumentFromQueue() throws GateException, IOException {
List<Hospital> hospitalList=clinicalDocumentRepository.getActiveHospitals();
for ( Hospital hospital: hospitalList) {
int hospitalId= hospital.getHospitalId();
log.info(hospitalId+"------------------------hospitalId");
List<Document> documents=clinicalDocumentService.getGateDocument(hospitalId);
for(Document document: documents) {
int documentId=0;
documentId=(int) document.getFeatures().get("documentId");
clinicalDocumentFactory.updateDocumentRunningStatus( documentId,2,hospitalId) ;
log.info("Processing document :"+document.getFeatures().get("documentId"));
try {
long analyisStartTime = System.currentTimeMillis();
documentProcessor.processDocument(document);
long analysisEndTime = System.currentTimeMillis();
long analysisDuration=(analysisEndTime-analyisStartTime);
log.info("analysis time{}",analysisDuration);
if(analysisDuration>60000)
log.warn("processing of this document is taking too long {}",documentId);
clinicalDocumentFactory.updateDocumentRunningStatus( documentId,1,hospitalId) ;
clinicalDocumentFactory.updateDocumentInitialRunningDate(documentId,hospitalId);
clinicalDocumentFactory.updateDocumentLastRunDate(documentId,hospitalId);
long insertionStartTime = System.currentTimeMillis();
annotationService.handleDocumentAnnotations(document ,hospitalId);
long insertionEndTime = System.currentTimeMillis();
long insertionDuration=(insertionEndTime-insertionStartTime);
log.info("insertion time{} ",insertionDuration);
if(insertionDuration>60000)
log.warn("Insertion of this document is taking too long {}",documentId);
saveDocToXml( document);
}
|
Cirs-gapp work flow